Database OrganizationThe release of the database includes 3 parts for each example: the raw measurement, the fitted BRDF parameters, and the STAF factorization. Due to the space limitation, we can not release all the 1,280 raw measurements. Instead, we selected 1 view and 6 lighting directions (thus 6 measurement in total) for each example at each time step. The structure of the database, for example, tvBTF09, is as follows:
Name ConventionThere are 26 samples in this database. Each sample has a unique id, from "tvBTF09" to "tvBTF45" (see here for a list of the samples). Take tvBTF09 as an example, the name convention of the database will be given as follows, step by step:
1) Texture Size where LLL is the light index (from 000 to 149), VV is the view index (from 00 to 15, see here for an explaination of the dome), and TT is the time index (form 00 to the time steps recorded for this sample). The image is in the EXR format. This is a format for HDR (high dynamic range) image. Please refer here for the information of this format. The raw data can be used in various ways. For example, in the BRDF fitting phase which will be explained right below, we do feel that neither the BRDF model we used nor the fitting method is perfect, especially for the specular parameters. One could try to use other types of BRDF models (e.g. Oren-Nayar + Full Torrance-Sparrow or Anisotropic Model), or other more robust fitting algorithm with these raw data. 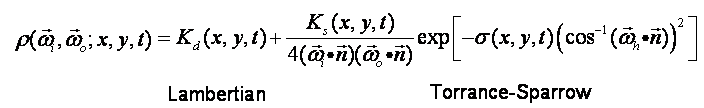 where there are 5 parameters: 3 for Kd (R,G,B), 1 for Ks, and 1 for Sigma. NOTE: Sigma defined in the above formula is different from the Sigma used in the original paper(Equation 1) for numerical reason. Their relationship is simple: Sigma_defined_here=1/(Sigma_in_paper)^2. So at each time step, we have 3 EXR images to represent these parameters:
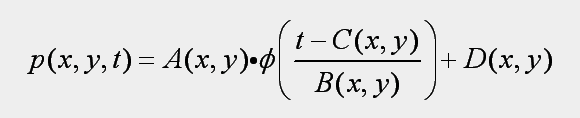 In this model, p(x,y,t) is one of the 5 BRDF parameters we have already fitted, and the variables in this model are the following 4 textures and 1 curve:
For each of the 5 BRDF parameters, we did the above STAF factorization. The results, for example, for the Kd parameters, are saved in the following files under this directory:
RedWe also included a figure to show these curves. For example, the R/G/B curves for Kd looks like this. Our implementation of the algorithm (in Matlab) to estimate the STAF model can be downloaded here (151KB). Note that there are few samples that can not get robust STAF factorization either for the specular BRDF parameters (Ks and Sigma), or for all the BRDF parameters The possible reasons are as follows:
Below is the list of these samples:
Image FormatImages in this database are saved in EXR format. EXR can be viewed as a float-point image type, used to represent HDR (High Dynamic Range) images. Compared with directly writing float point value to a raw file, EXR uses a "Half" type and has an internal compress algorithm to reduce the size of the image. More information about EXR format and library to read/write EXR images in C/C++ (and also a PhotonShop plugin) can be found at OpenEXR website. There is also a set of handy tools for EXR manipulation available here.In our study, I only used it to write a 3-channel float-point value image (for grey level images, the values in the 3 channels are the same), and did not use any fancy features in OpenEXR. I wrote two simple matlab functions to read/write EXR images for this simple usage. Download here (790 KB). Micah Kimo Johnson also wrote a very nice Matlab tool for read/write EXR images here. |
||||||||||||||||||||||||||||||
STAF Database Home Contact: staf@lists.cs.columbia.edu Last modified: 08/28/2006 |